DSCAM
DSCAM and Dscam are both abbreviations for Down syndrome cell adhesion molecule.[1] The case difference between the two abbreviations is significant as DSCAM refers to the gene that encodes for the Dscam protein.[2]
Down syndrome (DS), caused by trisomy 21, is the most common birth defect associated with intellectual disability. Recently, the novel gene DSCAM has been identified in the DS critical region. DSCAM is predicted to be a transmembrane protein with a very high structural and sequence homology to the immunoglobulin (Ig) superfamily of cell adhesion molecules. It is expressed in the developing nervous system with the highest level of expression occurring in the fetal brain. When this gene is over-expressed in the developing fetal central nervous system, it leads to Down syndrome. Diverse glycoproteins of cell surfaces and extracellular matrices, operationally termed as 'adhesion molecules' are important in the specification of cell interactions during development as well as maintenance and regeneration of the nervous system.[3]
A homologue of the Dscam protein in Drosophila melanogaster has 38,016 isoforms[4] arising from four variable exon clusters (12, 48, 33 and 2 alternatives, respectively).[1] By comparison, the entire Drosophila melanogaster genome only has 15,016 genes. The diversity of isoforms from alternative splicing of the Dscam1 gene in D. melanogaster allows every neuron in the fly to display a unique set of Dscam proteins on its cell surface. Dscam interaction stimulates self-avoidance mechanisms that are essential for normal neural circuit development.[5]
History/discovery
The DSCAM protein structure is conserved, with roughly more than 20% amino acid identity across the deuterostomes and protostomes, and assuming an ancestral homologous gene, places the origin of the DSCAM gene at >600 million years ago. Since then, the DSCAM gene has been duplicated at least once in vertebrates and insects.[6][7]
DSCAM was first identified in an effort to characterize proteins located within human chromosome band 21q22, a region known to play a critical role in Down syndrome.[8] The name Down syndrome cell adhesion molecule was chosen for a combination of reasons including: 1) chromosomal location, 2) its appropriate (normal) expression in developing neural tissue, and 3) its structure as an Ig receptor related to other CAMs.[9]
DSCAM gene
The DSCAM gene has been identified in the DS critical region. Dscam is predicted to be a transmembrane protein and a member of the immunoglobulin (Ig) superfamily of cell adhesion molecules. It is expressed in the developing nervous system with the highest level of expression occurring in the fetal brain. When this gene is over-expressed in the developing fetal central nervous system, it leads to Down syndrome. Diverse glycoproteins of cell surfaces and extracellular matrices, operationally termed as 'adhesion molecules' are important in the specification of cell interactions during development as well as maintenance and regeneration of the nervous system.[3]
Another DSCAM-like gene, DSCAML1, is located on chromosome band 11q23, a locus associated with Gilles de la Tourette and Jacobsen syndromes.[10]
Some intriguing changes in the gene structure of DSCAM have occurred in arthropods where several duplications of exons generated three large tandem arrays that are alternatively spliced.[11] This alternative splicing of individual exon sequences within an array occurs in a mutually exclusive and combinatorial manner allowing for expression of tens of thousands of Dscam isoforms. In the arthropods' genomes these three large exon arrays encode the N-terminal halves of the second and third Ig domains and the full Ig7 domain.[7][11][12][13]
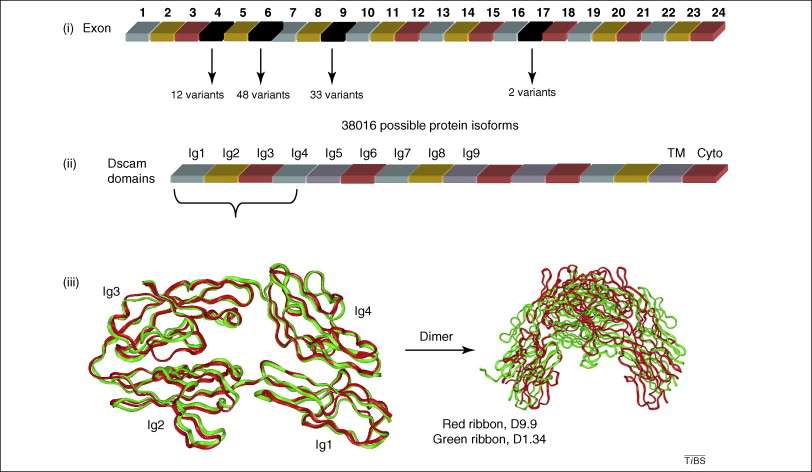
Alternative splicing generates large number of isomers with different protein–protein interaction possibilities. (i) Dscam has 24 exons; exon 4 has 12 variants, exon 6 has 48 variants, exon 9 has 33 variants, and exon 17 has two variants. The combination of exons 4, 6, and 9 leads to 19,008 possible isoforms with different extracellular domains (due to differences in Ig2, Ig3 and Ig4). With two different transmembrane domains from exon 17, the total possible protein products could reach 38,016 isoforms. (ii) These isoforms have different specific interactions due to structural variability. (iii) For example, the crystal structures of two isoforms (first four domains only) revealed large variation in their binding epitopes, resulting in totally different dimer forms. Red ribbons represent the structure of isoform D9.9 which is the product of variant 9 of exon 4 and variant 9 of exon 6 (PDB code 2V5R), the green ribbons represents isoform D1.34, which is the product of variant 1 of exon 4 and variant 34 of exon 6 (PDB code 2V5S). The figure in the lower left corner is the superimposition of the two monomers of D9.9 and D1.34 with the differences in the large Ig3 domain loop highlighted. The figure in the lower right corner is the superimposition of two dimers of D9.9 and D1.34, with a completely different interface and dimer conformations.[14]
Comparing the homology between genes and their products, is fundamental in understanding the phylogenetic relationship across the evolutionary pathway. In addition to the thousands of isoforms that can be populated from a single DSCAM of one species, DSCAM also demonstrates a diverse array of homology across species. Below are the genes, mRNA transcripts, and proteins identified as homologs of Down syndrome adhesion molecule.
| Species | Gene | mRNA | Protein |
|---|---|---|---|
| H.sapiens | DSCAM | NM_001389.3 | NP_001380.2 |
| P.troglodytes | DSCAM | XM_001171538.1 | XP_001171538.1 |
| M.mulatta | DSCAM | XM_002803124.1 | XP_002803170.1 |
| C.lupus | DSCAM | XM_544893.3 | XP_544893.3 |
| B.taurus | DSCAM | XM_002685111.2 | XP_002685157.1 |
| M.musculus | Dscam | NM_031174.4 | NP_112451.1 |
| R.norvegicus | Dscam | NM_133587.1 | NP_598271.1 |
| G.gallus | DSCAM | XM_416734.3 | XP_416734.3 |
| D.rerio | dscam | NM_001030224.1 | NP_001025395.1 |
| D.melanogaster | CG42330 | NM_001043131.2 | NP_001036596.2 |
| A.gambiae | AgaP_AGAP007092 | XM_308666.4 | XP_308666.4 |
Functions
Like many neuronal receptors, Dscam proteins have multiple functions, with repulsive and attractive roles that are dependent on the type of ligand that they interact with.
Immunity
Invertebrates do not have antibody-based immune systems. Instead, invertebrates rely on their innate immune system to eliminate infectious entities. The task of detecting and responding to a diverse pool of infectious agents are accomplished by germline encoded pattern recognition receptors (PRRs), which detect different patterns associated with the molecular markers to initiate an immune response.[15][16][17][18][19] The role of Dscam in the fly immune response was demonstrated by an RNAi mediated depletion experiment of DSCAM in which it was found to be associated with the cells that play a role in the fly's immune system.[15][20]
Dscam is found to have a role in phagocytosis in insects. The splicing pattern of the gene accompanying the phagocytic activity is specific to the type of infectious pathogen. In mosquitoes, the silencing of the Anopheles gambiae Dscam (AgDscam) disables its capacity to fight Plasmodium. The specificity of the Dscam recognition mechanism allows the mosquitoes of this species to differentiate the infection between bacteria and Plasmodium, and between Plasmodium berghei and Plasmodium falciparum.[15][21]
Regulation of synaptogenesis
Self-avoidance is a mechanism where the neuronal processes from the cell repel each other during arborization and axon branching to avoid fasciculation and clumping. Self-avoidance is necessary to prevent extensive overlapping in the arborization pattern and to facilitate the coverage of the neuronal processes across different regions of the nervous system during development.
DSCAM is recognized to be involved in this process in both vertebrates and invertebrates during neural development. Cell aggregation assays show that cell adhesion molecules, such as DSCAM, belonging to the immunoglobulin superfamily bind homophilically and specifically.[22][23][24][25] These molecules also appear to have a role in chemoattraction and repulsion.
Dscam1, of drosophila, may be one of the molecules involved in counteracting the netrin-dependent chemoattraction between neuronal processes during the neural development stage.[22] As previously mentioned, the Dscam1 gene in drosophila can encode 19008 extracellular domains, which bind homophilically and with isoform specificity.[26] The isoform-specific binding properties of Dscam, during homophilic repulsion, are the basis of self-avoidance, which is a crucial developmental mechanism for uniform distribution of axonal and dendritic processes in the formation of synaptic fields.[5] The neurons express a stochastic array of Dscam1 isoforms on their cell surface. Cells that have the same isoform patterns displaced on their surface, recognize the other as 'self', which leads to self-avoidance with the processes of neurons of the same subtype homophilically repelling from each other.
In addition to homophilic repulsion, Dscam1 mediates repulsion between neurites of different subtypes based on the specific isoform patterns displayed on the cell surface. This is called cell-type specific avoidance. The photoreceptor terminals of Drosophila form synapses with the postsynaptic invariant (tetrad) synapses that connect a pair of postsynaptic elements. Dscam is thought to aid this process by regulating the synaptic specificity through exclusion of inappropriate synaptic combination at the contact site.[27]
Furthermore, DSCAM is thought to have a role in 'tiling' during the drosophila's neuronal development. Tiling is a mechanism in which the processes from cells that share the same function work to create nerve bundles in a defined territory to create a pattern of non-overlapping dendritic or axonal fields.[28] Dscam1 and Dscam2 appear to be involved in axonal branching and tiling in Drosophila.[29][30] Tiling occurs when homophilic repulsion mediated by Dscam2 prevents the processes of the same class of cells from overlapping.[5] While both Dscam1 and Dscam2 mediate homophilic repulsion, the Dscam2 gene (unlike Dscam1) only encodes two alternative isoforms and thus lacks possible molecular diversity.[27] Consequently, the role of Dscam2, in either self-avoidance or cell-type-specific avoidance, occurs depending on which isoform or ratio of isoforms that the neuron expresses.[27]
Interactions
Many Ig superfamily molecules bind homophilically and heterophilically, and Dscam/DSCAM proteins are no exception. Vertebrate DSCAMs and DSCAML1s have not only been shown to bind homophilically (i.e., DSCAM–DSCAM or DSCAML1–DSCAML1, and not DSCAM–DSCAML1),[31][32] but also have cell-type specific, mutually exclusive, expression patterns.[32][33] Due to the combinatorial use of alternative exons, the homophilic binding specificity of Drosophila Dscam is amplified to tens of thousands of potential homodimers.,[34][35] Biochemical assays (cell-to-cell and bead-to-cell binding assays) were used to demonstrate that isoform-specific homodimerization occurs with remarkable binding specificity. This reveals that Dscam diversity can give rise to >18,000 distinct homodimers.[9]
Clinical significance
The role of Ig-CAMs in human development and disease is only beginning to be elucidated. This may be of particular interest with respect to the DSCAMs, as DSCAM maps to chromosome 21 in a region critical for the neurocognitive and other defects of Down syndrome[8][36] and DSCAML1 maps to chromosome 11 in a region whose deletion is associated with 11q deletion syndrome. This gives rise to neurocognitive defects and a subset of other defects which are similar to those seen in DS, including psychomotor retardation, Strabismus, Epicanthus, Telecanthus, carp-shaped upper lip, low-set dysmorphic ears, and cardiac defects.[33][37] The level of DSCAM expression is increased by more than 20% in the DS brain.[38] Given its identity as a potential neural morphogen and its expression in the cerebral and cerebellar cortices from the earliest stages in their development, it is not unreasonable to suggest that this level of DSCAM over-expression may contribute to the pre- and post-natal defects of DS, particularly, the cerebral and cerebellar hypoplasia and the abnormalities of the dendritic tree.[9][39] Further, a role for DSCAM over-expression in contributing to the defects of cortical lamination seen in DS[40] is supported by the fact that disruptions in other genes expressed by Cajal–Retzius cells, such as Reelin and LIS1, cause severe defects in neuroblast migration and cortical lamination.[41][42]
A study of congenital heart defect (CHD) investigated the polygenic effect of DSCAM with other genes. Under normal physiological conditions, DSCAM and COL6A2 work jointly in the drosophila to mediate cell matrix adhesion. However, over-expressing DSCAM and COL6A2 in the drosophila and mouse heart, resulted in a high mortality rate in addition to several serious heart defects, including atrial septal defects and cardiac hypertrophy. The interaction between DSCAM and COL6A2 and their combined effects were also observed in the H9c2 cardiac cell line with incidence of cardiac hypertrophy. While other gene combinations were screened to test the polygenic effect on the cardiac disorder, the DSCAM – COL6A2 pair was found to cause the most severe adverse effect in drosophila.[43] Translating the result to human cases of heart defects in DS patients require more study due to species-specific variance in the gene expression level. Nonetheless, the finding that DSCAM exerts a synergistic effect on the cardiac disease progression, upon disrupted expression level, allows future research on its role in some other major diseases.
See also
References
- 1 2 Schmucker D, Clemens JC, Shu H, Worby CA, Xiao J, Muda M, Dixon JE, Zipursky SL (June 2000). "Drosophila Dscam is an axon guidance receptor exhibiting extraordinary molecular diversity". Cell. 101 (6): 671–84. doi:10.1016/S0092-8674(00)80878-8. PMID 10892653.
- ↑ Alves-Sampaio, Alexandra; José Antonio Troca-Marín; María Luz Montesinos (6 October 2010). "NMDA-Mediated Regulation of DSCAM Dendritic Local Translation Is Lost in a Mouse Model of Down's Syndrome". The Journal of Neuroscience.
- 1 2 Agarwala KL, Nakamura S, Tsutsumi Y, Yamakawa K (2000). "Down syndrome cell adhesion molecule DSCAM mediates homophilic intercellular adhesion.". Brain Res Mol Brain Res. 79 (1–2): 118–26. doi:10.1016/s0169-328x(00)00108-x. PMID 10925149.
- ↑ Neves, G.; Zucker J; Daly M; Chess A. (February 2004). "Stochastic yet biased expression of multiple Dscam splice variants by individual cells.". Nature Genetics. 36 (3): 240–246. doi:10.1038/ng1299. PMID 14758360.
- 1 2 3 Hattori D, Millard SS, Wojtowicz WM, Zipursky SL (2008). "Dscam-mediated cell recognition regulates neural circuit formation". Annu. Rev. Cell Dev. Biol. 24 (1): 597–620. doi:10.1146/annurev.cellbio.24.110707.175250. PMC 2711549
 . PMID 18837673.
. PMID 18837673. - ↑ Crayton, MackE; Powell, BradfordC; Vision, ToddJ; Giddings, MorganC (2006). BMC Evolutionary Biology. 6 (1): 16. doi:10.1186/1471-2148-6-16. ISSN 1471-2148. Missing or empty
|title=(help)
- 1 2 Brites, D.; McTaggart, S.; Morris, K.; Anderson, J.; Thomas, K.; Colson, I.; Fabbro, T.; Little, T. J.; Ebert, D.; Du Pasquier, L. (2008). "The Dscam Homologue of the Crustacean Daphnia Is Diversified by Alternative Splicing Like in Insects". Molecular Biology and Evolution. 25 (7): 1429–1439. doi:10.1093/molbev/msn087. ISSN 0737-4038.
- 1 2 Yamakawa, K (1998). "DSCAM: a novel member of the immunoglobulin superfamily maps in a Down syndrome region and is involved in the development of the nervous system". Human Molecular Genetics. 7 (2): 227–237. doi:10.1093/hmg/7.2.227. ISSN 1460-2083.
- 1 2 3 Schmucker, D.; Chen, B. (2009). "Dscam and DSCAM: complex genes in simple animals, complex animals yet simple genes". Genes & Development. 23 (2): 147–156. doi:10.1101/gad.1752909. ISSN 0890-9369.
- ↑ Lal Agarwala, Kishan; Ganesh, Subramaniam; Tsutsumi, Yukie; Suzuki, Toshimitsu; Amano, Kenji; Yamakawa, Kazuhiro (2001). "Cloning and Functional Characterization of DSCAML1, a Novel DSCAM-like Cell Adhesion Molecule That Mediates Homophilic Intercellular Adhesion". Biochemical and Biophysical Research Communications. 285 (3): 760–772. doi:10.1006/bbrc.2001.5214. ISSN 0006-291X.
- 1 2 Schmucker, Dietmar; Clemens, James C; Shu, Huidy; Worby, Carolyn A; Xiao, Jian; Muda, Marco; Dixon, Jack E; Zipursky, S.Lawrence (2000). "Drosophila Dscam Is an Axon Guidance Receptor Exhibiting Extraordinary Molecular Diversity". Cell. 101 (6): 671–684. doi:10.1016/S0092-8674(00)80878-8. ISSN 0092-8674. PMID 10892653.
- ↑ Graveley, B. R. (2004). "The organization and evolution of the Dipteran and Hymenopteran Down syndrome cell adhesion molecule (Dscam) genes". RNA. 10 (10): 1499–1506. doi:10.1261/rna.7105504. ISSN 1355-8382.
- ↑ Watson, F. L. (2005). "Extensive Diversity of Ig-Superfamily Proteins in the Immune System of Insects". Science. 309 (5742): 1874–1878. doi:10.1126/science.1116887. ISSN 0036-8075. PMID 16109846.
- ↑ Tsai, Chung-Jung; Ma, Buyong; Nussinov, Ruth (2009). "Protein–protein interaction networks: how can a hub protein bind so many different partners?". Trends in Biochemical Sciences. 34 (12): 594–600. doi:10.1016/j.tibs.2009.07.007. ISSN 0968-0004.
- 1 2 3 Smith, Paul; Mwangi, J.; Afrane, Y.; Yan, G.; Obbard, D.; Randford-Cartwright, L.; Little, T. (2011). "Alternative Splicing of the Anopheles Gambiae Dscam gene in Diverse Plasmodium Falciparum Infections". Malaria Journal. 10: 156. doi:10.1186/1475-2875-10-156.
- ↑ R, Medzhitov; Janeway C (2000). "Innate immune recognition: mechanisms and pathways". Immunol. Rev. 173: 89–97. doi:10.1034/j.1600-065x.2000.917309.x.
- ↑ Christophides, GK; Vlachou D; Kafatos FC (2004). "Comparative and functional genomics of the innate immune system in the malaria vector Anopheles gambiae". Immunol Rev. 198: 127–148. doi:10.1111/j.0105-2896.2004.0127.x. PMID 15199960.
- ↑ Akira, S; Takeda K (2004). "Toll-like receptor signalling". Nat Rev Immunol. 4: 499–511. doi:10.1038/nri1391.
- ↑ Christensen, BM; Li JY; Chen CC; Nappi AJ (2005). "Melanization immune responses in mosquito vectors". Trends Parasitol. 21: 191–199.
- ↑ Watson, FL; Puttmann-Holgado R; Thomas F; Lamar DL; Hughes M; Kondo M; Rebel VI; Schmucker D (2005). "Extensive diversity of Ig-superfamily proteins in the immune system of insects". Science. 309 (5742): 1874–1878. doi:10.1126/science.1116887. PMID 16109846.
- ↑ Dong, YM; Taylor HE; Dimopoulos G (2006). "AgDscam, a hypervariable immunoglobulin domain-containing receptor of the Anopheles gambiae innate immune system". PLoS Biology. 4: 1137–1146. doi:10.1371/journal.pbio.0040229.

- 1 2 Garrett, Andrew M.; Tadenev, Abigail L. D.; Burgess, Robert W. (1 January 2012). "DSCAMs: restoring balance to developmental forces". Frontiers in Molecular Neuroscience. 5. 5. doi:10.3389/fnmol.2012.00086.
- ↑ Lal Agarwala, Kishan; Ganesh, Subramaniam; Tsutsumi, Yukie; Suzuki, Toshimitsu; Amano, Kenji; Yamakawa, Kazuhiro (1 July 2001). "Cloning and Functional Characterization of DSCAML1, a Novel DSCAM-like Cell Adhesion Molecule That Mediates Homophilic Intercellular Adhesion". Biochemical and Biophysical Research Communications. 285 (3): 760–772. doi:10.1006/bbrc.2001.5214.
- ↑ Agarwala, Kishan Lal; Nakamura, Sawako; Tsutsumi, Yukie; Yamakawa, Kazuhiro (1 June 2000). "Down syndrome cell adhesion molecule DSCAM mediates homophilic intercellular adhesion". Molecular Brain Research. 79 (1–2): 118–126. doi:10.1016/S0169-328X(00)00108-X.
- ↑ Yamagata, Masahito; Sanes, Joshua R. (24 January 2008). "Dscam and Sidekick proteins direct lamina-specific synaptic connections in vertebrate retina". Nature. 451 (7177): 465–469. doi:10.1038/nature06469.
- ↑ Wojtowicz, Woj M.; Flanagan, John J.; Millard, S.Sean; Zipursky, S.Lawrence; Clemens, James C. (1 September 2004). "Alternative Splicing of Drosophila Dscam Generates Axon Guidance Receptors that Exhibit Isoform-Specific Homophilic Binding". Cell. 118 (5): 619–633. doi:10.1016/j.cell.2004.08.021. PMC 2691713
 . PMID 15339666.
. PMID 15339666. - 1 2 3 Millard, S. Sean; Lu, Zhiyuan; Zipursky, S. Lawrence; Meinertzhagen, Ian A. (9 September 2010). "Drosophila Dscam Proteins Regulate Postsynaptic Specificity at Multiple-Contact Synapses". Neuron. 67 (5): 761–768. doi:10.1016/j.neuron.2010.08.030.
- ↑ Grueber, W. B.; Sagasti, A. (23 June 2010). "Self-avoidance and Tiling: Mechanisms of Dendrite and Axon Spacing". Cold Spring Harbor Perspectives in Biology. 2 (9): a001750–a001750. doi:10.1101/cshperspect.a001750.
- ↑ Wang, J; Zugates, CT; Liang, IH; Lee, CH; Lee, T (14 February 2002). "Drosophila Dscam is required for divergent segregation of sister branches and suppresses ectopic bifurcation of axons.". Neuron. 33 (4): 559–71. doi:10.1016/S0896-6273(02)00570-6. PMID 11856530.
- ↑ Millard, SS; Flanagan, JJ; Pappu, KS; Wu, W; Zipursky, SL (7 June 2007). "Dscam2 mediates axonal tiling in the Drosophila visual system.". Nature. 447 (7145): 720–4. doi:10.1038/nature05855. PMC 2691714
 . PMID 17554308.
. PMID 17554308. - ↑ Agarwala, Kishan Lal; Nakamura, Sawako; Tsutsumi, Yukie; Yamakawa, Kazuhiro (2000). "Down syndrome cell adhesion molecule DSCAM mediates homophilic intercellular adhesion". Molecular Brain Research. 79 (1–2): 118–126. doi:10.1016/S0169-328X(00)00108-X. ISSN 0169-328X.
- 1 2 Yamagata, Masahito; Sanes, Joshua R. (2008). "Dscam and Sidekick proteins direct lamina-specific synaptic connections in vertebrate retina". Nature. 451 (7177): 465–469. doi:10.1038/nature06469. ISSN 0028-0836.
- 1 2 Barlow, Gillian M; Micales, Bruce; Chen, Xiao-Ning; Lyons, Gary E; Korenberg, Julie R (2002). "Mammalian DSCAMs: roles in the development of the spinal cord, cortex, and cerebellum?". Biochemical and Biophysical Research Communications. 293 (3): 881–891. doi:10.1016/S0006-291X(02)00307-8. ISSN 0006-291X.
- ↑ Wojtowicz, Woj M.; Flanagan, John J.; Millard, S.Sean; Zipursky, S.Lawrence; Clemens, James C. (2004). "Alternative Splicing of Drosophila Dscam Generates Axon Guidance Receptors that Exhibit Isoform-Specific Homophilic Binding". Cell. 118 (5): 619–633. doi:10.1016/j.cell.2004.08.021. ISSN 0092-8674. PMC 2691713
 . PMID 15339666.
. PMID 15339666. - ↑ Wojtowicz, Woj M.; Wu, Wei; Andre, Ingemar; Qian, Bin; Baker, David; Zipursky, S. Lawrence (2007). "A Vast Repertoire of Dscam Binding Specificities Arises from Modular Interactions of Variable Ig Domains". Cell. 130 (6): 1134–1145. doi:10.1016/j.cell.2007.08.026. ISSN 0092-8674.
- ↑ Barlow, G.M.; Micales, B.; Lyons, G.E.; Korenberg, J.R. (2001). "Down Syndrome Cell Adhesion Molecule is conserved in mouse and highly expressed in the adult mouse brain". Cytogenetic and Genome Research. 94 (3–4): 155–162. doi:10.1159/000048808. ISSN 1424-859X.
- ↑ Epstein, C.J. (1986). "The Consequences of Chromosomal Imbalance". Cambridge University Press, Cambridge.
- ↑ Bahn, Sabine; Mimmack, Michael; Ryan, Margaret; Caldwell, Maeve A; Jauniaux, Eric; Starkey, Michael; Svendsen, Clive N; Emson, Piers (2002). "Neuronal target genes of the neuron-restrictive silencer factor in neurospheres derived from fetuses with Down's syndrome: a gene expression study". The Lancet. 359 (9303): 310–315. doi:10.1016/S0140-6736(02)07497-4. ISSN 0140-6736.
- ↑ Raz, N.; Torres, I. J.; Briggs, S. D.; Spencer, W. D.; Thornton, A. E.; Loken, W. J.; Gunning, F. M.; McQuain, J. D.; Driesen, N. R.; Acker, J. D. (1995). "Selective neuroanatornic abnormalities in Down's syndrome and their cognitive correlates: Evidence from MRI morphometry". Neurology. 45 (2): 356–366. doi:10.1212/WNL.45.2.356. ISSN 0028-3878.
- ↑ Golden JA, Hyman BT (1994). "Development of the superior temporal neocortex is anomalous in trisomy 21.". J Neuropathol Exp Neurol. 53 (5): 513–20. doi:10.1097/00005072-199409000-00011. PMID 8083693.
- ↑ Ogawa M, Miyata T, Nakajima K, Yagyu K, Seike M, Ikenaka K, et al. (1995). "The reeler gene-associated antigen on Cajal-Retzius neurons is a crucial molecule for laminar organization of cortical neurons.". Neuron. 14 (5): 899–912. doi:10.1016/0896-6273(95)90329-1. PMID 7748558.
- ↑ Noctor SC, Palmer SL, Hasling T, Juliano SL (1999). "Interference with the development of early generated neocortex results in disruption of radial glia and abnormal formation of neocortical layers.". Cereb Cortex. 9 (2): 121–36. doi:10.1093/cercor/9.2.121. PMID 10220225.
- ↑ Grossman, Tamar R.; Gamliel, Amir; Wessells, Robert J.; Taghli-Lamallem, Ouarda; Jepsen, Kristen; Ocorr, Karen; Korenberg, Julie R.; Peterson, Kirk L.; Rosenfeld, Michael G.; Bodmer, Rolf; Bier, Ethan; Barsh, Gregory S. (3 November 2011). "Over-Expression of DSCAM and COL6A2 Cooperatively Generates Congenital Heart Defects". PLoS Genetics. 7 (11): e1002344. doi:10.1371/journal.pgen.1002344.

Additional sources
- Li W, Guan KL (July 2004). "The Down syndrome cell adhesion molecule (DSCAM) interacts with and activates Pak". J. Biol. Chem. 279 (31): 32824–31. doi:10.1074/jbc.M401878200. PMID 15169762.
- Wojtowicz WM, Flanagan JJ, Millard SS, Zipursky SL, Clemens JC (September 2004). "Alternative splicing of Drosophila Dscam generates axon guidance receptors that exhibit isoform-specific homophilic binding". Cell. 118 (5): 619–33. doi:10.1016/j.cell.2004.08.021. PMC 2691713
 . PMID 15339666.
. PMID 15339666. - Watson FL, Püttmann-Holgado R, Thomas F, et al. (September 2005). "Extensive diversity of Ig-superfamily proteins in the immune system of insects". Science. 309 (5742): 1874–8. doi:10.1126/science.1116887. PMID 16109846.
- Chen BE, Kondo M, Garnier A, et al. (May 2006). "The molecular diversity of Dscam is functionally required for neuronal wiring specificity in Drosophila". Cell. 125 (3): 607–20. doi:10.1016/j.cell.2006.03.034. PMID 16678102.