Molecular recognition
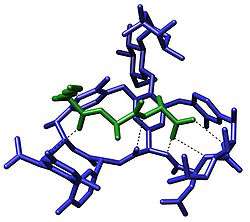
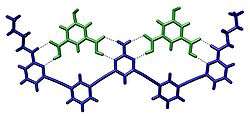

The term molecular recognition refers to the specific interaction between two or more molecules through noncovalent bonding such as hydrogen bonding, metal coordination, hydrophobic forces,[3][4] van der Waals forces, π-π interactions, halogen bonding, electrostatic and/or electromagnetic[5] effects. In addition to these direct interactions as well solvent can play a dominant indirect role in driving molecular recognition in solution.[6][7] The host and guest involved in molecular recognition exhibit molecular complementarity.[8][9]
Biological systems
Molecular recognition plays an important role in biological systems and is observed in between receptor-ligand,[10][11] antigen-antibody, DNA-protein, sugar-lectin, RNA-ribosome, etc. An important example of molecular recognition is the antibiotic vancomycin that selectively binds with the peptides with terminal D-alanyl-D-alanine in bacterial cells through five hydrogen bonds. The vancomycin is lethal to the bacteria since once it has bound to these particular peptides they are unable to be used to construct the bacteria’s cell wall.
Synthetic molecular recognition
Recent work suggests that molecular recognition elements can be synthetically produced at the nano-scale,[12] circumventing the need for naturally-occurring molecular recognition elements for the development of sensing tools for small molecules. Bio-mimetic polymers such as peptoids can be used to recognize larger biological targets such as proteins [13] and the conjugation of polymers to synthetic fluorescent nanomaterials can generate synthetic macromolecular structures that serve as synthetic antibodies for optical protein recognition and detection.[14]
Supramolecular systems
Chemists have demonstrated that artificial supramolecular systems can be designed that exhibit molecular recognition. One of the earliest examples of such a system are crown ethers which are capable of selectively binding specific cations. However, a number of artificial systems have since been established.
Static vs. dynamic
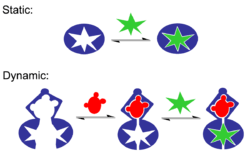
Molecular recognition can be subdivided into static molecular recognition and dynamic molecular recognition. Static molecular recognition is likened to the interaction between a key and a keyhole; it is a 1:1 type complexation reaction between a host molecule and a guest molecule to form a host-guest complex. To achieve advanced static molecular recognition, it is necessary to make recognition sites that are specific for guest molecules.
In the case of dynamic molecular recognition the binding of the first guest to the first binding site of a host affects the association constant of a second guest with a second binding site.[15] In the case of positive allosteric systems the binding of the first guest increases the association constant of the second guest. While for negative allosteric systems the binding of the first guest decreases the association constant with the second. The dynamic nature of this type of molecular recognition is particularly important since it provides a mechanism to regulate binding in biological systems. Dynamic molecular recognition may enhance the ability to discriminate between several competing targets via the conformational proofreading mechanism. Dynamic molecular recognition is also being studied for application in highly functional chemical sensors and molecular devices.
Complexity in molecular recognition
A recent study based on molecular simulations and compliance constants describes molecular recognition as a phenomenon of organisation. Even for small molecules like carbohydrates, the recognition process can not be predicted or designed even assuming that each individual hydrogen bond's strength is exactly known.[16] However, as Mobley et al.[17] concluded, the accurate prediction of the molecular recognition events needs to go beyond the static snapshot of a single frame between the guest and the host. Entropies are key contributors to binding thermodynamics and need to be accounted for in order to predict more accurately the recognition process. Entropies are not observable in single bound structures (static snapshot).
See also
References
- ↑ Knox, James R.; Pratt, R. F. (July 1990). "Different modes of vancomycin and D-alanyl-D-alanine peptidase binding to cell wall peptide and a possible role for the vancomycin resistance protein" (Free full text). Antimicrobial Agents and Chemotherapy. 34 (7): 1342–7. doi:10.1128/AAC.34.7.1342. PMC 175978
 . PMID 2386365.
. PMID 2386365. - ↑ Bielawski, Christopher; Chen, Yuan-Shek; Zhang, Peng; Prest, Peggy-Jean; Moore, Jeffrey S. (1998). "A modular approach to constructing multi-site receptors for isophthalic acid" (Free full text). Chemical Communications (12): 1313–4. doi:10.1039/a707262g.
- ↑ "The Binding of Benzoarylsulfonamide Ligands to Human Carbonic Anhydrase is Insensitive to Formal Fluorination of the Ligand". Angew. Chem. Int. Ed. 52 (30): 7714–7717. 2003. doi:10.1002/anie.201301813. PMID 23788494.
- ↑ "Water Networks Contribute to Enthalpy/Entropy Compensation in Protein–Ligand Binding". J. Am. Chem. Soc. 135 (41): 15579–15584. 2013. doi:10.1021/ja4075776. PMID 24044696.
- ↑ Cosic, I (1994). "Macromolecular bioactivity: is it resonant interaction between macromolecules?—theory and applications". IEEE transactions on bio-medical engineering. 41 (12): 1101–14. doi:10.1109/10.335859. PMID 7851912.
- ↑ Baron, Riccardo; Setny, Piotr; McCammon, J. Andrew (2010). "Water in Cavity-Ligand Recognition". Journal of the American Chemical Society. 132 (34): 12091–12097. doi:10.1021/ja1050082. PMC 2933114
 . PMID 20695475.
. PMID 20695475. - ↑ Baron, Riccardo; McCammon, J. Andrew (2013). "Molecular Recognition and Ligand Binding". Annual Review of Physical Chemistry. 64: 151–175. doi:10.1146/annurev-physchem-040412-110047. PMID 23473376.
- ↑ Lehn, Jean-Marie (1995). Supramolecular Chemistry. Weinheim: Wiley-VCH. ISBN 978-3-527-29312-4. OCLC 315928178.
- ↑ Gellman, Samuel H. (1997). "Introduction: Molecular Recognition". Chemical Reviews. 97 (5): 1231–1232. doi:10.1021/cr970328j. PMID 11851448.
- ↑ "The Binding of Benzoarylsulfonamide Ligands to Human Carbonic Anhydrase is Insensitive to Formal Fluorination of the Ligand". Angew. Chem. Int. Ed. 52 (30): 7714–7717. 2003. doi:10.1002/anie.201301813. PMID 23788494.
- ↑ "Water Networks Contribute to Enthalpy/Entropy Compensation in Protein–Ligand Binding". J. Am. Chem. Soc. 135 (41): 15579–15584. 2013. doi:10.1021/ja4075776. PMID 24044696.
- ↑ Zhang, Landry, Strano et al. Nature Nanotechnology 2013 http://www.nature.com/nnano/journal/v8/n12/full/nnano.2013.236.html
- ↑ Mannige, Ranjan V.; Haxton, Thomas K.; Proulx, Caroline; Robertson, Ellen J.; Battigelli, Alessia; Butterfoss, Glenn L.; Zuckermann, Ronald N.; Whitelam, Stephen (2015-10-15). "Peptoid nanosheets exhibit a new secondary-structure motif". Nature. 526 (7573): 415–420. doi:10.1038/nature15363. ISSN 0028-0836.
- ↑ Beyene, Abraham G.; Demirer, Gozde S.; Landry, Markita P. (2009-01-01). Current Protocols in Chemical Biology. John Wiley & Sons, Inc. doi:10.1002/cpch.10/full. ISBN 9780470559277.
- ↑ Shinkai, Seiji; Ikeda, Masato; Sugasaki, Atsushi; Takeuchi, Masayuki (2001). "Positive allosteric systems designed on dynamic supramolecular scaffolds: toward switching and amplification of guest affinity and selectivity". Accounts of Chemical Research. 34 (6): 494–503. doi:10.1021/ar000177y. PMID 11412086.
- ↑ Complexity in Molecular Recognition, Phys. Chem. Chem. Phys., 2011, 13, 10136-10146
- ↑ Binding of Small-Molecule Ligands to Proteins: "What You See" Is Not Always "What You Get", Structure 17(4): 489-498, 2009
External links
- http://www.mdpi.org/ijms/sections/molecular-recognition.htm Special Issue on Molecular Recognition in the Int. J. Mol. Sci.